In this next instalment of our series of articles looking at patenting cannabis-related inventions, our focus is on the growth in industrial biotechnology in this area: genetically modifying plants or host organisms to produce key intermediates in the cannabinoid pathway, or end cannabinoids themselves.
Historically, as commercial and medical interest in cannabinoids increased, technologies, and resulting patent filings, focussed on methodologies for extracting relevant cannabinoids from cannabis plant extracts. Such techniques are inherently limited by the quantities of active compounds, or their precursors, naturally produced by the plants: supply of sufficient levels of plant materials has to keep up with the demand, while being constrained by the requirement to only grow plants in a secure environment, and the large quantities of energy and water required to grow the plants and isolate the cannabinoid products.
In a world concerned with climate change and energy demand, interest has therefore naturally turned to more industrial processes, in particular, the use of genetic engineering to increase enzymatic turnover within the cannabinoid pathway, either in cannabis plants for increased yield after extraction, or in host systems such as E.coli or yeast.
Exploiting the cannabinoid pathway
The biosynthesis of cannabinoids in nature is shown in Figure 1, and includes the enzymes olivetolic acid cyclase and prenyltransferase, to make the key intermediate cannabigerolic acid (CBGA). CBGA is then converted by cannabinoid synthases to either Δ9-THCA or CBDA, intermediates which are non-enzymatically converted to Δ9-tetrahydrocannabinol (the psychoactive component of cannabis) and cannabidiol respectively.
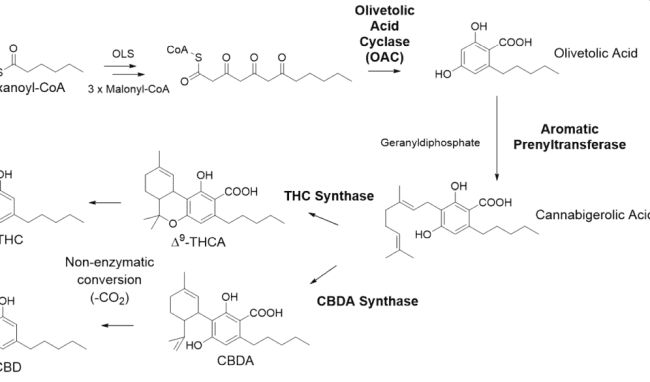
Figure 1: Cannabinoid Pathway
Companies such as Genomatica and Ginkgo Bioworks are developing extensive patent portfolios around variant enzymes along these pathways, and engineered cells which express these enzymes. In recent years, Genomatica’s WO2020/214951, WO2020/247741, WO2021/211611, and WO2021/046367 have published, directed to olivetol synthase variants, olivetolic acid cyclase variants, cannabinoid synthase variants, and prenyltransferase variants respectively.
Ginkgo Bioworks’ recent publications include WO2021/195520 (directed to variant terminal synthases), (WO2021/257915 (directed to olivetolic acid cyclase variants), WO2022/011175 (directed to variant terminal synthases, and WO2022/081615 (directed to prenyltransferase variants and related chimeric prenyltransferases and fusion polypeptides).
Protecting variants
With so much patenting activity in this area, notably from US companies, what do patent applicants need to think about when preparing patent applications, to maximise their chances of obtaining broad, commercially useful protection in Europe?
The most cost-effective way of obtaining broad geographical coverage is of course via a European patent application at the European Patent Office (EPO). However, a little forethought and planning before filing a US provisional application, and/or a PCT application, can increase the chances of obtaining allowance of claims with a usefully broad scope in Europe.
While there is a basic requirement at the EPO that a gene sequence is only patentable if it is capable of industrial application, this is easily met for most cases – particularly in the cannabinoid sector – in which there is a recognised therapeutic application requiring industrial production of active compounds: a simple statement in the specification setting out the industrial applicability usually suffices.
Patent claims in the field of genetically modified organisms are typically directed to polypeptide sequences of variant enzymes, and corresponding nucleotide sequences of the genes that encode the variant enzymes, with the claims defined by reference to one or more specific sequences disclosed in a sequence listing filed as part of or with the patent application. Examples of such claims are:
A nucleic acid having a sequence that is at least 80% identical to SEQ ID NO:1 and wherein the nucleic acid encodes a polypeptide having olivetolic acid cyclase activity.
A polypeptide comprising a sequence that is at least 80% identical / similar to SEQ ID NO:2.
A polypeptide including at least 80% homology to SEQ ID NO:2.
Sequence identity – what does it even mean?
“Sequence identity” refers to the number or percentage of nucleotides/amino acids that match exactly between two different sequences over a defined length in a given alignment, while “sequence similarity” refers to a resemblance between two sequences when compared.
Based on the open language of the above example claims (“having”, “comprising”, “including”), the sequence identity can be determined in different ways, depending, for example, on whether the subject and query sequences have the same length or different lengths, or how the sequences are aligned. How this is defined in the claims and specification can lead to very different results when it comes to assessing novelty of such a claim. In addition, the EPO may also consider such claims unclear and thus not allowable.
Figure 2 shows why it is advisable, if not essential, to specify the algorithm used, or at the very least how the identity is to be defined:
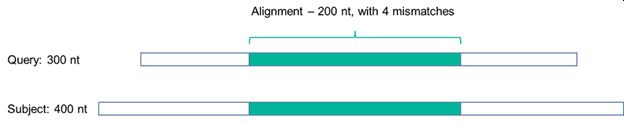
Figure 2: A sequence alignment
Looking at the hypothetical example in Figure 2, if the identity is defined with reference just to the alignment over the 200 nt sequence, then the percentage identity is calculated as being 98% (with the calculation being (200-4)/200 = 98%).
However, if the identity is defined with reference to a 300 nt query sequence (e.g., the sequence inputted into the algorithm), then there are 104 mismatches, and the % identity over the query drops to 65%. Further still, if the identity is defined with reference to a 400 nt subject sequence (i.e., hits within the algorithm sequence database), then there are 204 mismatches and the % identity over the subject sequence drops further to 49%.
Algorithms such as FastA and BLAST provide identity over the best local alignment, whereas GLSearch provides identity over the query sequence, GSSearch provides identity over the subject sequence and GGsearch identity over a global alignment.
The EPO’s Guidelines for Examination at F-IV, 4.24 explain how “sequence identity” and “sequence similarity” are interpreted, and unequivocally state that if no algorithm, calculation method or similarity-scoring matrix is provided or defined in the application, the broadest interpretation of these terms will be applied, using any reasonable algorithm, calculation method or matrix known at the relevant filing date.
In addition, if the similarity or homology is the only feature to distinguish the subject-matter of a claim from the prior art, the EPO will raise a clarity objection unless the determination or calculation of the percentage of homology is clearly defined in the application as filed.
Using the above example, loss of identity between what a patent applicant intends (for example BLAST local alignment) and what the EPO might search (alignment over a larger subject sequence) could result in a variant sequence being considered to lack novelty and/or be unclear if no algorithm was defined. As a result, without suitable fallback positions (e.g., to the algorithm, or the length of sequence) included at the drafting stage it may not be possible to amend the claims to restore novelty.
Engineered or mutant enzymes that have higher yields, faster turnover, or higher enantioselectivity are sought after in any industrial setting and are commodities or assets that need to be protected. It is important that patent applicants do not lose sight of the European Patent Office’s examination practices for biotechnological inventions defined by nucleotide or amino acid sequence (or, indeed, the entry into force on 1 July 2022 of ST26 relating to the preparation of sequence listings – see here) if a competitive edge in Europe is to be secured.
The content of this article is intended to provide a general guide to the subject matter. Specialist advice should be sought about your specific circumstances.